SNOMED Is A Knowledge Graph
The Systematized Nomenclature of Medicine, Clinical Terms (SNOMED CT) is a de facto standard for standardizing clinical vocabulary and ontology in many parts of the world, including in Singapore. You can access SNOMED CT in a number of ways. If you work in a large healthcare organization, it probably has a subscription that you can apply for access through. For the rest of us, you can sign up for free with the United States’ National Library of Medicine (NLM). The NLM maintains the Unified Medical Language System (UMLS), a behemoth chimaera of biomedical vocabularies that include SNOMED. With a NLM account, you can download SNOMED from the UMLS site. The International Edition is what you want if working on SNOMED in Singapore. Do note that there are conditions and restrictions of use which you should comply with. There is also a publicly accessible online SNOMED CT browser at https://browser.ihtsdotools.org .
The SNOMED ontology framework comprises 3 atomic parts: concepts, descriptions and relationships. A SNOMED concept is a unique, machine-readable clinical idea represented by an identification number (the Concept ID). Concepts are linked to one another via relationships, in hierarchical parent-child fashion as well as laterally across concept domains. A concept can have more than one parent.
Descriptions are human-language phrases that describe concepts. Naturally there are often multiple descriptions of a given concept, so one canonical description is chosen to be the “Fully Specified Name (FSN)” of the concept. The other descriptions then become essentially synonyms of the FSN.
Relationships link concepts. There are 2 main relationship types: the “IS A” relationship type links a child concept to its parent concept (eg “Viral Pneumonia” IS A “Pneumonia”), and attribute relationships link concepts from different hierarchies (eg “Finding Site” relationship links “Pain” (a “Symptom”) to “Abdomen” (a “Body Part”). Architecturally, SNOMED CT ontology is a knowledge graph, and can be instantiated as a graph database (GDB).
Language Models and Graph Databases
Language models interact with GDBs in a number of ways. Two valuable ones are: (1) during the construction of a knowledge graph, and (2) during graph-based retrieval-augmented generation (GraphRAG).
When building a knowledge graph from text, a common step is to use a large language model (LLM) to extract entities and their relationships from the text, often as a source -> relationship -> destination triplet. For example, you could extract symptoms and examination findings related to a disease condition in a clinic consultation. LLMs typically excel at this sort of language interpretation tasks. By extension therefore, LLMs can also assist in schema design and ontology engineering.
The graph database then becomes an essential data infrastructure if you had a need for GraphRAG in your use case. Indications for GraphRAG in healthcare might include
- explainable and traceable reasoning paths required for fostering trust and auditability,
- catering for evolving information sources, such during the course of civil emergencies (hours to days), disease outbreaks (days to months) and longitudinal lifelong patient health records (years and decades),
- Synthesizing information from disparate information sources, e.g. linking cancer knowledgebases with patient registries.
GDBs are a natural fit for the relationship-rich text in healthcare prose, as well as the multi-hop traversals in healthcare networks and records. Languages, especially coding languages, are essentially graphs. So theoretically, there should be synergy in GDBs leveraging on LLMs for healthcare. In practice, LLMs tend to be more proficient in SQL than graph query languages like Cypher, due to the larger codebase of SQL available for LLM training. Relational databases are also a more ‘operationally mature’ ecosystem, with greater support (SQL linters, query optimizers, debugging tools, established patterns etc….). Thus, there is intriguing research opportunity to explore the use of LLMs with GDBs for healthcare, possibly sparking creative adjacent-possibles that are difficult to achieve in the relational space.
Building a SNOMED Concept Database
To start off exploration, I built a simple SNOMED concept database in Neo4j on my Windows desktop PC. Neo4j is a widely used graph database with strong community support, which is the main reason for choosing it for experimentation. Other popular graph databases include MemGraph, TigerGraph and ArangoDB.
This is a minimalistic database that is a static snapshot of concepts, descriptions and relationships. Each concept or description node has just enough attributes (referred to as “properties” in Neo4j) to map relationships; several properties from the original SNOMED downloads were not modelled as they were not felt to be useful.
I document the steps here for those of you who would like to try it as well. The project ran reasonably stably on an old i7 SkyLake PC with 32Gb RAM on Windows, YMMV. There are 3 parts to this exercise:
- Download, clean and prep the SNOMED CT International Version files from NLM. NLM updates SNOMED versions twice yearly. I used the December 2025 version.
- Install Neo4j Desktop. You can also try using the free cloud-based AuraDB service by Neo4j or the open-source Neo4j Community Edition, but the rest of this blog is based on Neo4j Desktop Version 2. If you’re using the Neo4j Community Edition, be prepared for a lot more tinkering with configurations, settings and plugin management.
- Load prepared SNOMED CT concepts, descriptions and relationships csv files into Neo4j.
Prepping SNOMED CT Files
SNOMED CT files come in two flavors – Full and Snapshot. We’ll be using the Snapshot version for this exploration. The Full version would eventually be required in a production system as it contains revisions such as retired concepts and ontology changes, which will necessarily need to be tracked in an ongoing production deployment. The required files are found in the Terminology subfolder of the download. There are three of them (‘XXXXXXXX’ refers to the date of release):
- sct2_Concept_Snapshot_INT_20XXXXXX.txt
- sct2_Description_Snapshot-en_INT_20XXXXXX.txt
- sct2_Relationship_Snapshot_INT_20XXXXXX.txt
I renamed them to concept.txt, description.txt and relationship.txt for ease of typing. The data preparation Python script is saved on my GitHub (https://github.com/eukairos/snomed). Of the outputs, we only need to use the snomed_concept.csv, snomed_description.csv and snomed_relationship.csv files.
Installing Neo4j Desktop Version 2
Installing Neo4j Desktop is straightforward. After installation, when prompted, key in a name for your instance, enter a password, and connect to the instance. The instance should contain 2 databases, ‘system’ and ‘neo4j’. You should only interact with the neo4j database; leave the system database untouched. You can’t rename the databases, unfortunately. Before populating the database, we should install a useful plugin: APOC (“Awesome Procedures On Cypher”). To do so, while you are still in the instance page (Click “Local instances” on the left menu – ie the menu on your left – if you’ve navigated away from the instance page), click on the hamburger menu at the top right, select “Plugins” and install the APOC plugin. Then restart the instance to effect the plugin.
Loading SNOMED Files
Neo4j uses a graph query language, Cypher, that is analogous to SQL for relational databases. The Cypher script for loading the files is saved on my GitHub (link as above). To load the files, follow the script carefully, especially the instructions for locating your Neo4j import directory.
You may also need to edit the Neo4j configuration file to increase Java heap size if your PC has small amount of RAM, see this video for how to do it, otherwise memory is auto-allocated usually.
To load the script, go into the Query page in Neo4j Desktop. This is found in the left menu under “Tools”. Once in the Query page, the Cypher editor is found at the right half of the screen. The topmost cell of the editor is empty – this is where you’ll cut-and-paste the script and run it. The run icon is at the right-hand end of the cell. Don’t run the entire script at one go, run it section by section – each section in the script is prefixed with one or more comments. It is similar to running a Jupyter notebook or an SQL query editor.
It may take several minutes to load the files, depending on your PC specs, because SNOMED files are large. Cypher has a propensity to silently fail from bugs in the script without alerting you, so sometimes it is a test of patience…
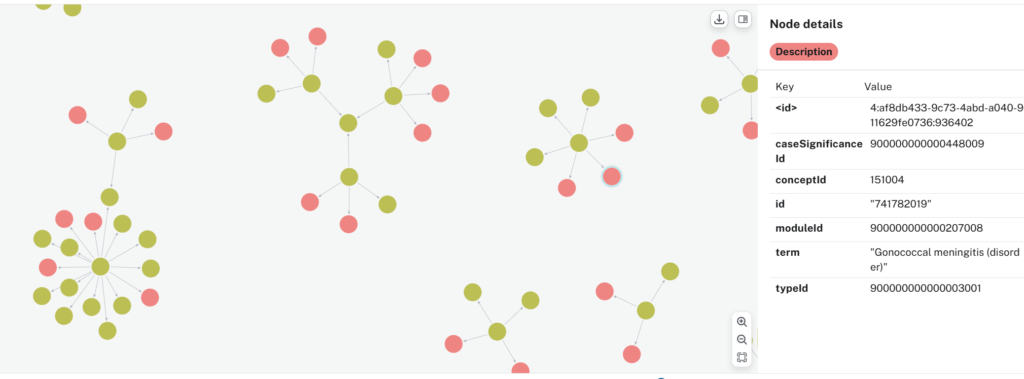
If all goes well, your output should be a SNOMED knowledge graph with:
- Unique concept nodes,
- Unique description nodes in many-to-one relationships with concept nodes,
- Concept nodes hierarchically linked via IS_A relationship,
- Concept nodes laterally linked via attribute relationships, the most common being “FINDING_SITE”.
Closing Remarks on Cypher Loading Script
During loading of the concept database, I used LMArena to assist with Cypher vibe coding, as I am new to Cypher. LMArena has a “Battle” mode in which two unidentified LLMs are pitted against each other. This was great for learning Cypher on-the-job as the two duelling LLMs often pointed out different aspects of Cypher in their generated responses. If you feedback on which one was the better performer, LMArena will reveal what it is. One of the more consistently good performers was Anthropic’s Claude Sonnet 4.5, but several others were also good. I also found (this is just a personal impression) that on the whole, they were not as accurate in Cypher compared to Python, and used deprecated Cypher statements quite often. Fortunately, the Cypher editor in Neo4j often is able to point out errors and suggest corrections.
If you want to take the opportunity to learn some basic Cypher, you could do the same, and develop the loading scripts yourself. If it works better than mine, send a PR, and I’ll upload it alongside mine.
This post assumes the reader has some basic Python knowledge.