In our previous graph database exercise, we built a graph of MIMIC-IV patients, their admissions, and diagnoses associated with each admission. In this exercise, we’ll load some of their allergies. The allergies documented in MIMIC-IV are not in some structured fields, but exist as free text inside clinical notes, which makes it a challenge to extract them out cleanly.
For this exercise, we’ll be using a derived dataset of MIMIC-IV that contains just notes; MIMIC-IV-EXT-BHC. The creators of this dataset have processed the original free text notes into sections; it’s much easier for us to work with this dataset than with the original notes. Patient allergies are in a section prefixed with <ALLERGIES>, so we can just extract that section out. But even so, the work is quite involved, and I haven’t actually completed it.
RxNorm
Before we start, you should know something about RxNorm, the drug dictionary I mapped these allergies to. RxNorm is the US National Library of Medicine’s standard drug dictionary, which is updated frequently. I used RxNorm because MIMIC-IV was assembled from US hospitals. In Singapore, we have the Singapore Standard Drug Dictionary (SDD) if you are working on a Singapore dataset. Unfortunately, the SDD is not released as a dataset, but in the form of an online formulary. You could try asking for access to the dataset from Synapxe.
The data model for RxNorm is rather complicated, but for our purpose here, we just need to know that, like SNOMED, RxNorm’s atomic unit is a concept, uniquely identified by an identifier “RXCUI”. Not all drug names make their way into an RXCUI though. The act of being entered into an RXCUI is called “normalization” (this term has abnormally different meanings in different contexts, here is yet another context….). So there will be some “unnormalized” drugs.
Unlike SNOMED concepts, RXCUI are not that fully ‘atomic’. Each RXCUI has within it RXAUI (for “atom unique identifiers”). These RXAUI are essentially synonyms of the same concept. So “Naproxen” has the RXCUI “198013” and each normalized name obtained from source vocabularies of naproxen (eg “Naproxen 250mg ORAL TABLET”, “naproxen@250 mg@ORAL@TABLET”) exist as RXAUIs linked to the RXCUI.
I decided to map the allergies to just 2 levels of concepts: the drugs themselves, and families of drugs, since this is how they are most commonly entered in structured data fields in health records eg “Bactrim” vs “Sulfonamides”.
Data Preparation
I divided up the cleaning and extracting into 3 Jupyter notebooks, all found on my GitHub. All 3 were done with Claude Sonnet’s help. The first notebook, “MIMIC_IV_Compile_Allergies.ipynb” breaks up the MIMIC-IV-EXT-BHC clinical notes into individual sections and save these individual sections for subsequent use. A subset of these sections, containing the contents of the <ALLERGIES> section of the original notes, was saved separately and used in the subsequent notebooks.
The 2nd, “MIMIC_IV_EXT_BHC_Normalize_Drugs_For_Allergies Part 1.ipynb” takes the saved entries, and maps them to RxNorm codes at either the name level or group level. For example, an allergy to “omeprazole” is mapped to the actual RxNorm code, while an entry for “penicillins” was mapped to the group level. Because not all drugs are normalized into RXCUIs, we end up with three levels of ‘certainty’ – those with RXCUIs mapped, those mapped at the drug level (Level 1 in my dataset) and those at the group level (Level 2 in my dataset).
The 3rd, “MIMIC_IV_EXT_BHC_Normalize_Drugs_For_Allergies Part 2.ipynb” splits out the subset of <ALLERGIES> entries that are matched to names in RxNorm into a .csv file, which I renamed ‘allergies.csv’ for easier typing. These are the ones that we’ll upload to our MIMIC graph. The unmatched ones I have stored in a separate .csv file – they’re quite messy, a mix of unrelated clinical notes, vital signs, social history etc…. all stored in the wrong section (or maybe, parsed wrongly by the 1st notebook). Some of them do contain actual allergies within their text, but I could not easily auto-extract them. We’ll have to use some other method, like entity-recognition and relationship linking, to extract them, if I ever get down to it.
Loading into Neo4j
Instead of using a Cypher script to load the allergies.csv file as we have done previously, I used the Neo4j Desktop UI to do so. It’s actually quite easy, just go to “Import” on the left hand menu of Neo4j Desktop, browse to my “allergies.csv” location, click on the middle pane to bring up the properties that I want on the right hand menu, and click “Map from file” on the right hand menu (See the figure below)
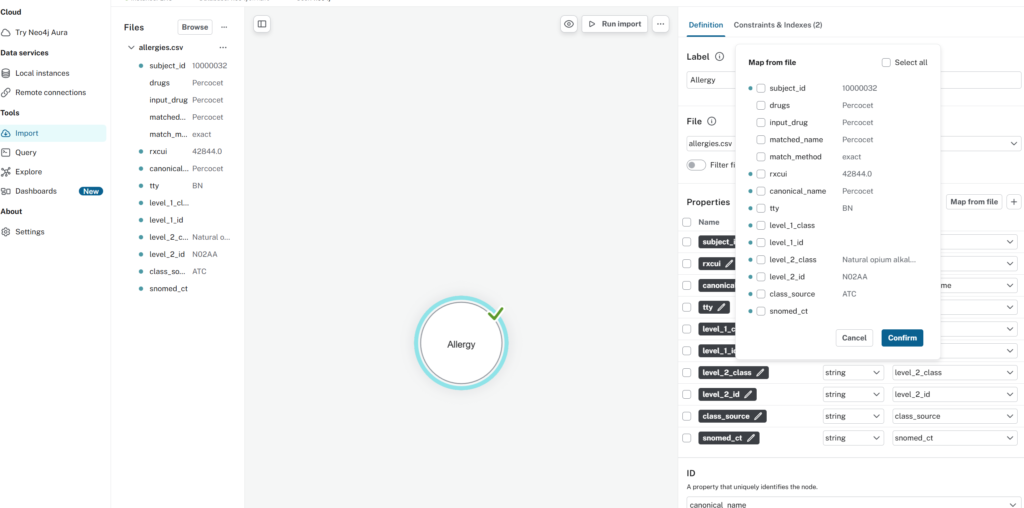
After that, I needed to
- Set up the relationship between patients and allergies,
- A bit of final cleaning for properties that should be strings, but were converted to floats, using the script below (also found on my GitHub – link above):

Here’s a view of some of the output:-
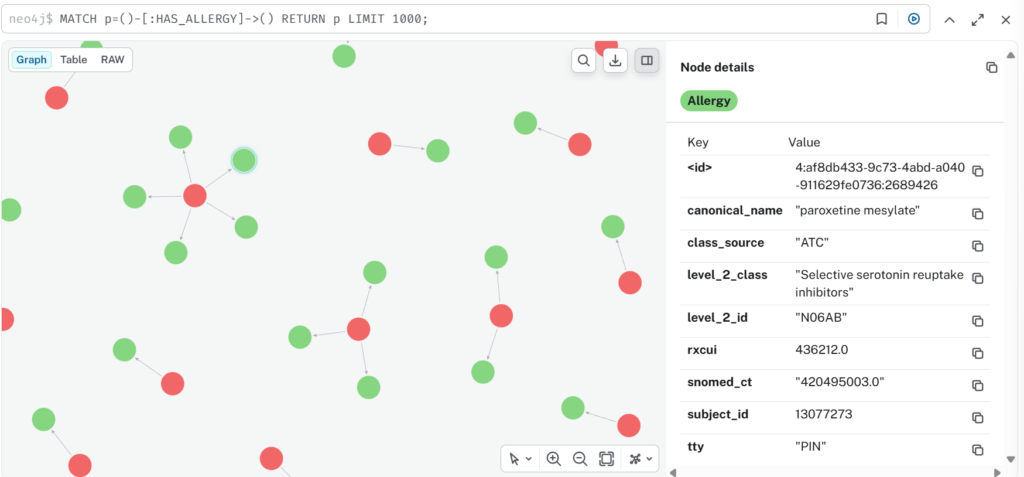
Closing Thoughts
This has been a more difficult task than I had anticipated for two reasons: (1) the data is in free text, and had lots of noise, (2) the RxNorm dictionary was new to me. Vibe coding with Claude helped save the day, and I learnt a lot from studying Claude’s code. The work is still unfinished, as I haven’t tackled the entries that were not mappable in this first effort.
If you are doing this for production, make sure you review carefully all the entries in your output files – allergy information can be a life-and-death difference. In my proof-of-concept project, I haven’t tackled production issues as yet.
Citations
MIMIC-IV-EXT-BHC: Aali, A., Van Veen, D., Arefeen, Y., Hom, J., Bluethgen, C., Reis, E. P., Gatidis, S., Clifford, N., Daws, J., Tehrani, A., Kim, J., & Chaudhari, A. (2025). MIMIC-IV-Ext-BHC: Labeled Clinical Notes Dataset for Hospital Course Summarization (version 1.2.0). PhysioNet. RRID:SCR_007345. https://doi.org/10.13026/5gte-bv70
PhysioNet: Goldberger, A., Amaral, L., Glass, L., Hausdorff, J., Ivanov, P. C., Mark, R., … & Stanley, H. E. (2000). PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation [Online]. 101 (23), pp. e215–e220. RRID:SCR_007345.
Leave a Reply