Clinical notes are peculiarly messy. Written under time pressure by busy clinicians, they’re full of abbreviations, shorthand, and — inevitably — typos. When you’re building natural language processing (NLP) pipelines that depend on these notes, these ‘features’ become a real problem. This post describes a tool Anthropic’s Claude helped me build to tackle that problem, and how you might adapt the same approach for other domains, or any other specialised domain.
The Problem: Invisible Errors in “Clean” Training Data
If you’re training a spell-correction model for clinical text, a common approach (following methods like NeuSpell and CIM) is to take a clean corpus, deliberately corrupt random words, and train the model to recover the originals. Simple enough — until you realise that your “clean” corpus isn’t actually clean.
Clinical discharge summaries have estimated error rates of 0.1–0.4% of tokens (1). That doesn’t sound like much, but across millions of tokens it amounts to thousands of mislabelled training examples. When the model correctly fixes a pre-existing typo like colection → collection, it gets penalised for deviating from the ground truth. Over time, this biases the model toward reproducing common clinical typos rather than correcting them.
I needed a systematic way to find and fix these pre-existing errors before they contaminated my training pipeline.
The Solution: Dictionary + AI + Human Review
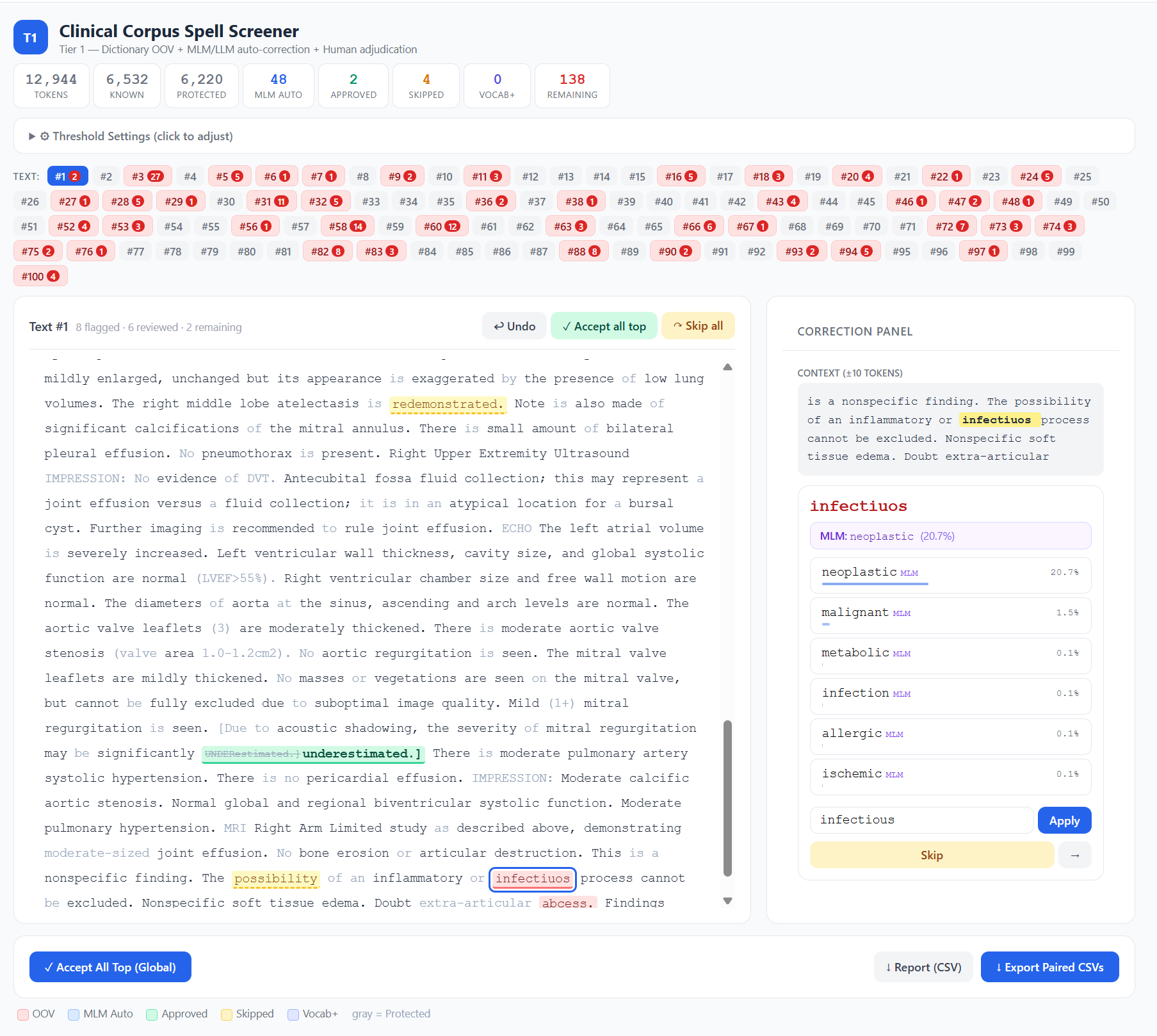
The Clinical Spell Screener is an open-source, browser-based tool that combines three layers of screening.
Layer 1: Dictionary-based OOV detection. Every token is checked against a 3-million-entry medical dictionary built from the UMLS Metathesaurus — covering SNOMED CT, RxNorm, LOINC, MeSH, ICD-10, and 20+ other clinical vocabularies — plus a curated list of 483 clinical abbreviations generated by Claude. Tokens absent from both are flagged as out-of-vocabulary (OOV). Crucially, guard patterns protect tokens that look unusual but are perfectly valid in clinical context: lab values like WBC-17.4*, abbreviation-colon patterns like HR:, and short tokens that are almost always legitimate codes.
Layer 2: Spell Correction. Flagged tokens are optionally evaluated by either edit distance, BioClinical-ModernBERT, a domain-specific masked language model (MLM), or an Ollama-based generative model. A MLM masks the flagged token in context and reads off the model’s softmax probability for each candidate correction. This gives you a real, calibrated probability — not a heuristic. A two-gate system prevents overcorrection: the candidate must exceed both an absolute confidence threshold and a ratio threshold relative to the original token’s probability. The model has to be confident and substantially more confident than it is in the original word.
Layer 3: Human-in-the-loop review. The browser interface shows colour-coded tokens — red for OOV, blue for model auto-corrections, green for human-approved — with a context window, candidate list, undo, and batch operations. Click a flagged token, choose a correction (or skip it, or add it to your vocabulary), and the cursor advances to the next one. When you’re done, the tool exports paired CSVs: the original text untouched, and the corrected version, ready for use as clean/noisy training pairs.
Why Not Just Use a Generative LLM?
It’s a fair question. I initially considered routing every OOV token through MedGemma or a similar generative model via Ollama. But the task — “is this word misspelled, and if so, what’s the correct word?” — is fundamentally a contextual word prediction problem, not a generation problem. A masked language model does this natively in a single forward pass. It’s faster (roughly 50×), produces calibrated probabilities rather than heuristic scores, and doesn’t require prompt engineering to avoid verbose explanations when you just need one word.
The tool still supports Ollama as a fallback for users who prefer it, but the MLM approach proved to be the better fit for this specific task.
In initial testing, I found that edit distance was sufficient, and often provided closer choices than MLM. It is also faster, so I have made it the default option of the tool.
Adapting It for Your Domain
Although I built this tool for clinical text, the architecture is domain-agnostic. If you work with legal documents, scientific papers, engineering reports, or any other specialised corpus, you can adapt it with three changes.
Replace the dictionary. The clinical dictionary is built from UMLS, but the tool accepts any plain text wordlist — one word per line. For legal text, you might extract terms from Black’s Law Dictionary or a legal ontology. For scientific text, use domain-specific vocabularies from your field. For general English, a standard wordlist like the one from en_US Hunspell combined with your domain’s terminology would work well.
Swap the language model. The tool accepts any HuggingFace masked language model via the --model flag. For general English, try bert-base-uncased or ModernBERT-base. For biomedical text, BioBERT or PubMedBERT are strong choices. For legal text, models like legal-bert-base-uncased exist. The key requirement is that it must be a masked LM (not a generative model) — that’s what gives you the calibrated probabilities.
Adapt the guard patterns. This is the part that requires the most thought. The clinical guard patterns are tuned for a specific kind of text: they protect tokens containing digits (lab values), colon-separated codes (HR:, BP:), slash-separated expressions ('w/o' (‘without’), 's/p' (‘status post’), and very short tokens (single or double characters that are usually valid abbreviations). If your domain has similar patterns — case numbers in legal text, chemical formulae in scientific text, part codes in engineering reports — you’ll want to define equivalent guards. The patterns live in the isProtected() function in index.html and are straightforward regex rules. The principle is simple: if a token pattern is almost always valid in your domain, protect it from being flagged, even if it’s not in your dictionary. Getting this right dramatically reduces false positives and makes the review process much more efficient.
The tool runs entirely locally — your data never leaves your machine — and the dictionary-only mode requires nothing more than Python and a browser. No GPU, no model download, no cloud API.
Try It
The tool is available on GitHub: github.com/eukairos/clinical-spell-checker
On Windows, download the repository, double-click start_lite.bat (dictionary-only) or start.bat (with AI assistance), and you’re running. On Mac/Linux, use start.sh or start_lite.sh. Upload your dictionary, abbreviations file, and corpus, and start reviewing.
If you use it in your research, I’d appreciate a citation — and if you adapt it for a new domain, I’d love to hear how it goes.
(1) Zhou L, Blackley SV, Kowalski L, Doan R, Acker WW, Landman AB, Kontrient E, Mack D, Meteer M, Bates DW, Goss FR. Analysis of Errors in Dictated Clinical Documents Assisted by Speech Recognition Software and Professional Transcriptionists. JAMA Netw Open. 2018 Jul;1(3):e180530. doi: 10.1001/jamanetworkopen.2018.0530. Epub 2018 Jul 6. PMID: 30370424; PMCID: PMC6203313.
This post was mostly written by Claude.
Leave a Reply