A Simple Patient Graph Database.
In a previous post, I shared how to build a SNOMED concept graph database. In this post, we will use the MIMIC-IV dataset (details below) to construct a graph of patients, their admissions and the diagnoses associated with each admission. We then link this graph to our SNOMED concept graph by matching diagnosis codes with SNOMED concept IDs.
Why would we want to do this? In principle, you can extend this concept to practically all the data available in MIMIC-IV, thus creating an electronic health record graph database, which would be the ultimate goal. For a start, we’ll work on something relatively simpler – just the diagnoses. Our purpose for this graph is to be able to find diagnoses and aggregate related diagnoses rapidly, such as all admissions related to tuberculosis, a disease that can surface at multiple body locations and in patients admitted to a wide range of clinical services.
There are several parts to this exercise:
- Download the relevant MIMIC-IV files.
- Download tables from the United States’ National Library of Medicine (NLM) for mapping ICD9CM and ICD10CM diagnosis codes to SNOMED.
- Prep the MIMIC-IV csv files. The main task here is to translate MIMIC diagnosis codes, which are coded in ICD9 or ICD10, into SNOMED concept IDs using the mapping tables downloaded.
- Load the processed MIMIC-IV patient, admission and diagnosis files into Neo4j, and mapping the MIMIC-IV diagnoses to their SNOMED concept IDs.
Download the relevant MIMIC-IV files
The starting point is the MIMIC-IV database, which is maintained at www.physionet.org. MIMIC stands for “Medical Information Mart for Intensive Care”, but it is, in fact, a comprehensive electronic health record dataset that includes also emergency department visits and hospital discharge summaries. You can read about it here: https://www.nature.com/articles/s41597-022-01899-x. PhysioNet grants credentialed access to researchers. The credentialing steps are detailed on MIMIC-IV’s webpage at https://physionet.org/content/mimiciv/3.1/ . Once credentialed, you may download the dataset. For our purposes, we’ll be using these files:
- patients.csv.gz
- services.csv.gz
- diagnoses_icd.csv.gz
- admissions.csv.gz
Download the ICD-to-SNOMED mapping tables
You can download the ICD9CM-to-SNOMED mapping tables, using the NLM account you created in the last post, from: https://www.nlm.nih.gov/research/umls/mapping_projects/icd9cm_to_snomedct.html. There are actually 2 tables: a one-to-one table (suffixed with ‘1TO1’) for codes with one-to-one matches, and a one-to-many (suffixed with ‘1TOM’). We’ll need both. ICD9CM stands for International Classification of Diseases 9 – Clinical Modification, which is slightly different from the ICD9 found in the MIMIC dataset. The differences are minor enough for us to accept for our exercise. For a production system that might reference these codes for, say, billing purposes, you’ll probably need a mapping source that is both accurate and maintainable.
The ICD10CM to SNOMED mapping table is found at the SNOMED US Edition page: https://www.nlm.nih.gov/healthit/snomedct/us_edition.html . The file we want is: tls_Icd10cmHumanReadableMap_US1000124_20250901.tsv (your version may be a later version). Again, ICD10CM (CM : Clinical Modification) is slightly different from original ICD10, but the differences are minor enough for us to accept.
Prep the MIMIC files
The Jupyter notebook for prepping the files, ‘Data Prep for loading MIMIC to Neo4j.ipynb’, can be found in my GitHub at https://github.com/eukairos/snomed/tree/main. Briefly, we’ll do the following:
- On the MIMIC files:
- Drop unused attributes from the source files.
- Reformat ICD9 and ICD10 codes into their standard formats.
- On the ICD-to-SNOMED mapping tables
- Drop unused attributes
- Concatenate the 3 files – ICD9CM one-to-one, ICD9CM one-to-many, and ICD10CM files into 1 mapping file.
- On the MIMIC diagnosis file, we’ll add in the new mapped SNOMED Concept IDs.
Our outputs are:
- A mimic_patients_age_gender.csv file that contains unique patient IDs, age and gender.
- A mimic_admissions_small.csv file that contains patient IDs, unique hospital admission IDs, admission date & time, and discharge date and time.
- A mimic_diagnoses_small.csv file that contains patient IDs, hospital admission IDs, diagnoses encoded in ICD and SNOMED.
- A mimic_disciplines.csv file that contains patient IDs, hospital admission IDs, transfer times, and current service admitted to. We’ll only use the hospital admission IDs and current service for this exercise, though.
Loading the MIMIC files
We take a pause here to review our schema for the graph. We will model the graph as:
- Patient nodes, with properties: age, gender, subject_id (the patient ID), and indexed on subject_id.
- Admission nodes, with properties: subject_id, hadm_id: the ID number of the admission, and also its index, admit time, and discharge time.
- Diagnosis nodes, which are tied to admissions, and have the properties:
- hadm_id: the admission episode to which a diagnosis is tied,
- snomed_cid: the SNOMED concept ID
4. Service nodes, which are the clinical services for each admission episode, with the properties:
- hadm_id: the admission episode for that patient’s stay with the clinical service, to which the service is tied,
- service: the string encoding of the clinical service.
This figure shows the overall schema including both MIMIC and SNOMED components.
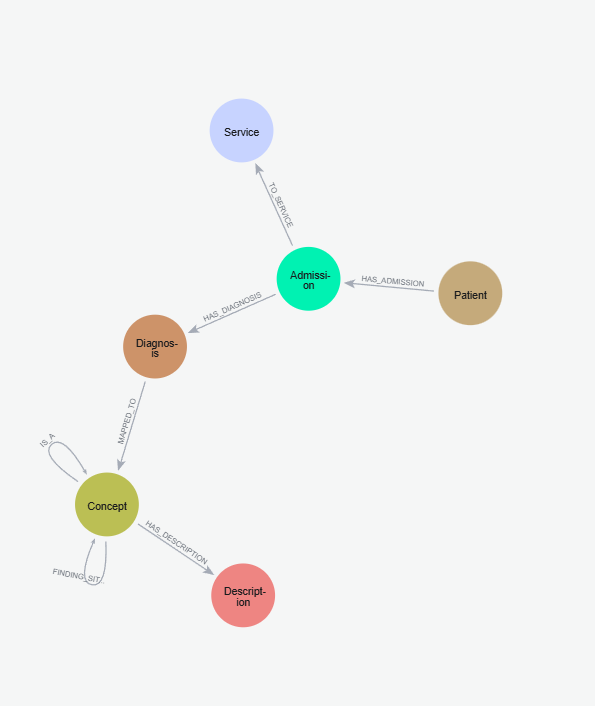
Based on our schema decisions above, we write the Cypher script to load the csv files into our Neo4j database that we set up in the last exercise. At the same time, we’ll link the diagnoses through their SNOMED IDs to our SNOMED Concept database. The script is also found in my GitHub (link above). You’ll need the APOC plugin installed, so do install it if you haven’t done so in the previous exercise. You should end up with a graph of patient nodes, admission nodes, diagnosis nodes and services nodes, linked to our SNOMED concept graph through the diagnosis nodes.
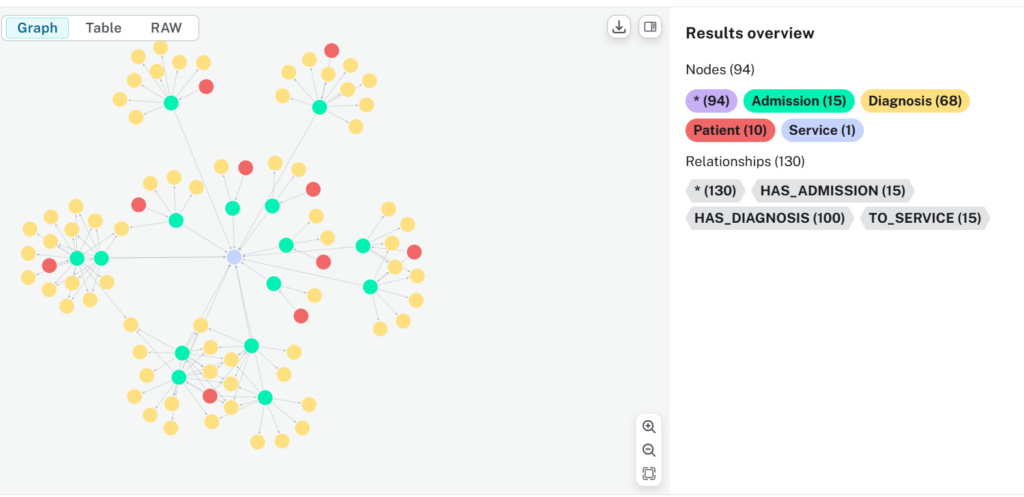
Putting Our Graph To Work
Let’s test our graph by trying to retrieve patients who have diagnoses related to “tuberculosis”, and have 1 or more hops to the search term “Tuberculosis (disorder)”. The Cypher script below can be found in my GitHub (link above).
| MATCH (search:Concept {id: ‘56717001’}) // tuberculosis, don’t forget the quotes WITH search MATCH (p:Patient)-[:HAS_ADMISSION]->(a:Admission)-[:HAS_DIAGNOSIS]->(d:Diagnosis)-[:MAPPED_TO]->(c:Concept)-[:HAS_DESCRIPTION]->(desc:Description) MATCH path=(c)-[:IS_A*1..]->(search) RETURN DISTINCT p.subject_id AS Patient_ID, desc.term AS Condition, length(path) AS Depth, [n IN nodes(path) | n.id] AS Hierarchy LIMIT 1000 |
- The first line of the script stores the concept ID for tuberculosis in a variable named ‘search’.
- The second line uses that concept ID as a filter.
- The third line of the script traces a path from Patient nodes, through Admission nodes, Diagnosis nodes, Concept nodes and finally Description nodes.
- The fourth line traces a path from children concept nodes that are at least one hop away from tuberculosis (ie the concept “tuberculosis” itself is not included) – that is the meaning of “[IS_A*1..]” in the line.
- The fifth line returns the variables that we want to see, and the last line limits output to 1000 lines.
This retrieves the table below
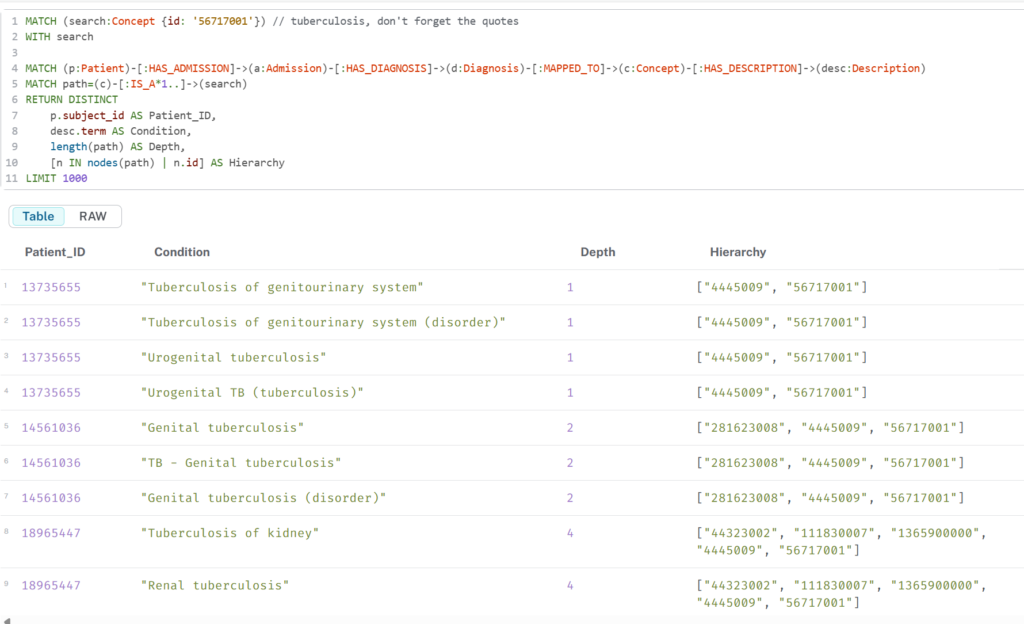
Limitations
You may have realized that our MIMIC-IV graph is just a snapshot of the MIMIC-IV data, and even then not a very accurate one. There are four main limitations to consider if you wish to build further on this graph:
- the patients’ ages are a snapshot at an “anchor age”. However, a patient may have multiple admissions over several years, and this is not modelled in the graph. In a production system, this would likely not be a problem as you likely have the birthdates of your patients and can track their ages across admissions.
- The ICD9 and ICD10 mapping tables to SNOMED are not a one-to-one. They are, in truth, many-to-many. In order to keep the database size manageable, for each unique ICD9 and ICD10 code, I took only the first SNOMED concept ID matched for this exercise. In addition, there are some ICD codes that have no mapped SNOMED concept IDs, these were dropped. So there is some information lost, hopefully not too much. If you really need all the codes to be mapped, the Jupyter notebook mentioned above does output the full versions.
- In the MIMIC databases, diagnoses have a sequence, indicating the “importance” of the diagnosis. A sequence number of 1 indicates it is the primary diagnosis for that admission episode. This information was not brought over to our graph. It should be relatively straightforward to incorporate this information, though, should you have a need to do so.
- For each admission, only the current service was included; this is the current service at discharge. In a single admission, there could be multiple transfers of a patient among several clinical disciplines. This information is found in another MIMIC-IV table that we did not use.
Closing Thoughts On Schema Design
You may have wondered why I linked the MIMIC-IV patient graph directly to the SNOMED Concept ID graph through the diagnoses. Perhaps a more maintainable schema might be to set up an interfacing mapping table instead, and maintain the 2 graphs separately, which you would need to do in a production system. That’s certainly viable, and your schema design will depend a lot on what you intend to do with your graphs. In our case, our aim was to be able to leverage graph traversal to arrive at summary diagnoses quickly, which this schema design allows.
Citations & Licensing Conditions
MIMIC-IV dataset: Johnson, A., Bulgarelli, L., Pollard, T., Gow, B., Moody, B., Horng, S., Celi, L. A., & Mark, R. (2024). MIMIC-IV (version 3.1). PhysioNet. RRID:SCR_007345. https://doi.org/10.13026/kpb9-mt58
Original MIMIC-IV publication: https://doi.org/10.1038/s41597-022-01899-x
PhysioNet citation: Goldberger, A., Amaral, L., Glass, L., Hausdorff, J., Ivanov, P. C., Mark, R., … & Stanley, H. E. (2000). PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation [Online]. 101 (23), pp. e215–e220. RRID:SCR_007345.
Licence: Any MIMIC-IV derived datasets or models should be treated as containing sensitive information. If you want to share PhysioNet data, you should share them on PhysioNet under the same agreement as the source data.
This post assumes the reader has some basic Python knowledge.